Pancreatic ductal adenocarcinoma (PDAC) is one of the clearest examples of why modern translational research needs more than literature synthesis. The biology is concentrated around a dominant oncogene, KRAS, but the clinical question is not simply whether KRAS is mutated. Scientists, clinicians, and drug developers need to know which allele is present, how that allele varies across cohorts, which drugs can engage it, what evidence supports the mechanism, what patient populations are still excluded, and how the trial landscape is moving.
That is the kind of question K-Dense Web is designed to answer as a research system rather than as a chat interface.
In one autonomous session, K-Dense Web built a reproducible analysis of the global RAS mutation landscape in pancreatic cancer. It queried the cBioPortal REST API, incorporated literature-derived cohorts for underrepresented geographies, stratified KRAS, NRAS, and HRAS mutations by allele and continent, computed drug-class eligibility, ran cross-continent statistical tests, generated publication-quality figures, mapped the RAS-targeted clinical-trial landscape from ClinicalTrials.gov, and produced a 19-page tumor-board technical brief.
The session ID was session_20260512_103421_e632f388fbf3. The scientific question was specific:
Characterize the prevalence of RAS gene mutations (KRAS, NRAS, HRAS) in pancreatic cancer cohorts worldwide, stratified by specific allele (G12D, G12V, G12C, G12R, Q61X, etc.) and by continent of cohort origin.
The output was not a generic summary. It was a structured research package with scripts, data tables, logs, figures, statistical tests, a trial landscape, and a peer-reviewed final brief.
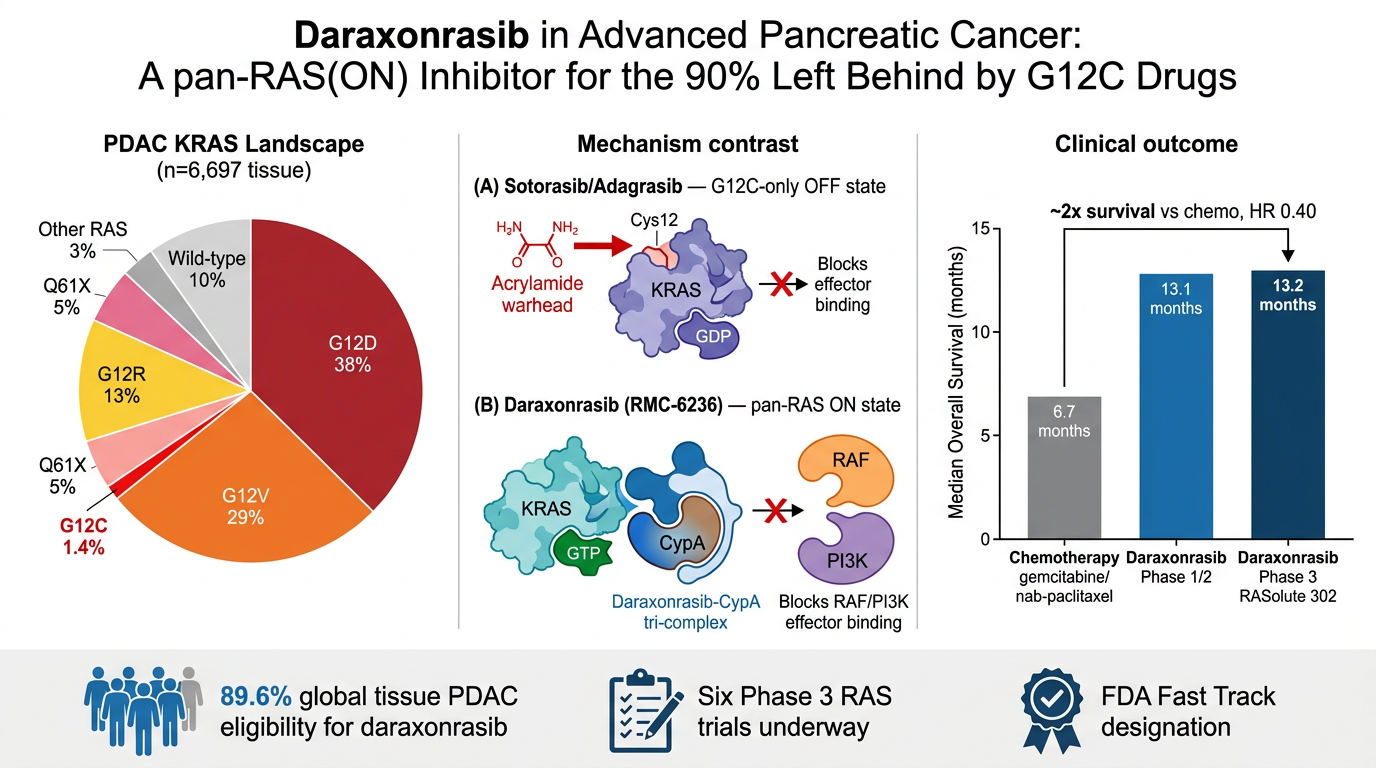
Why This Question Matters
RAS biology has moved from "undruggable" to one of the most active areas in oncology drug development. The first clinical breakthrough came from covalent KRAS-G12C inhibitors such as sotorasib and adagrasib. Those drugs validated KRAS as a therapeutic target, but their chemistry is allele-specific: they require the cysteine residue created by the G12C substitution.
That is a major limitation in PDAC. KRAS-G12C is common enough in lung adenocarcinoma to anchor a development program, but it is rare in pancreatic cancer. In PDAC, the dominant alleles are G12D, G12V, and G12R. A researcher evaluating a new RAS-directed therapy therefore needs more than a binary KRAS-mutant versus KRAS-wild-type classification. The relevant unit is allele-level prevalence, with enough cohort provenance to know whether a finding is robust across geography, tumor type, and assay modality.
This is especially important for pan-RAS(ON) inhibitors such as daraxonrasib (RMC-6236), which are designed to engage active, GTP-bound RAS through a cyclophilin A mediated tri-complex. Unlike G12C covalent inhibitors, this mechanism does not depend on the mutant cysteine at codon 12. In principle, it can cover a much broader set of activating RAS alleles.
The key translational question becomes quantitative:
How many PDAC patients are plausibly addressable by a pan-RAS(ON) mechanism compared with G12C-only inhibition, and does that conclusion hold across continents?
The Workflow
K-Dense Web decomposed the problem into five linked research tasks.
- Genomic data acquisition and stratification. Query cBioPortal for curated pancreatic-cancer studies, parse RAS-family protein changes into canonical hotspot alleles, add literature cohorts where public database coverage is sparse, and aggregate results by continent.
- Population overlap and eligibility analysis. Classify observed alleles into drug-coverage classes: G12C-only, pan-RAS(ON)-addressable, uncovered RAS variants, and RAS-wild-type.
- Visualization. Generate figures suitable for a scientific or tumor-board audience, including allele heatmaps, stacked allele distributions, eligibility comparisons, and a mechanism schematic.
- Statistical comparison. Test whether allele frequencies differ across continents after restricting to tissue-based PDAC cohorts and correcting for multiple testing.
- Clinical-trial landscape. Query ClinicalTrials.gov for RAS-targeted pancreatic and advanced solid tumor trials, annotate mechanisms, classify monotherapy versus combination strategies, and identify pivotal programs.
The workflow produced CSV, JSON, PDF, PNG, and LaTeX outputs. The important point is not that K-Dense summarized public knowledge. It created a traceable computational path from API calls to figures to written interpretation.
Data Sources and Cohort Construction
The genomic analysis used two complementary sources.
First, K-Dense Web queried 21 curated pancreatic cancer studies from the cBioPortal REST API. For each study, it retrieved the mutation molecular profile, the sequenced-sample list, and bulk mutation records for KRAS, NRAS, and HRAS. Protein-change strings were parsed into canonical alleles, including G12D, G12V, G12C, G12R, G12A, G12S, G13D, Q61H, Q61R, Q61L, Q61K, and A146 variants.
Second, K-Dense Web added six literature-derived cohorts to improve coverage for regions that are underrepresented in public genomic portals. These included Japanese, Chinese, German, Mediterranean, and Latin American PDAC cohorts. Each contributing study was tagged by tumor type so PDAC could be separated from pancreatic neuroendocrine tumors, acinar cell carcinoma, pancreatoblastoma, and mixed cohorts.
That tumor-type separation matters. PDAC has high-frequency KRAS hotspot biology. PNET and acinar tumors do not. Pooling them would dilute the biological signal and produce a misleading denominator.
The primary analytic cohort for eligibility and statistics was therefore tissue-based PDAC, not all pancreatic tumors and not all assay modalities. For the statistical comparison, K-Dense Web explicitly excluded a large MSK ctDNA cohort because lower liquid-biopsy sensitivity could deflate apparent mutation frequencies in North America. That left an apples-to-apples tissue-PDAC denominator of 6,697 samples:
| Continent | Tissue PDAC samples |
|---|---|
| Asia | 1,025 |
| Europe | 1,678 |
| North America | 3,390 |
| Oceania | 482 |
| South America | 122 |
This is the kind of methodological choice that determines whether a downstream conclusion is credible. A general model might say "KRAS is common in PDAC." A reproducible research workflow needs to say which cohorts were included, which assay types were excluded, why the denominator changed, and how the decision affects the result.
The Mutation Landscape: G12D, G12V, and G12R Dominate
Across continents, KRAS mutation prevalence in PDAC was high and consistent with the known biology of the disease. In the broader PDAC aggregation, KRAS-mutant prevalence ranged from 83.9% to 91.2% across continents. After isolating tissue cohorts for the eligibility analysis, North American prevalence rose into alignment with other regions, confirming that the lower pooled North American estimate was largely driven by ctDNA sensitivity rather than a major biological difference.
The allele-level result was more important than the overall KRAS rate. G12D was the most common KRAS allele across all five continents, representing roughly 34% to 45% of all PDAC tumors. G12V followed at about 27% to 30%, and G12R accounted for about 11% to 16%. G12C, the only allele directly covered by sotorasib and adagrasib, remained rare at roughly 1% to 2%.
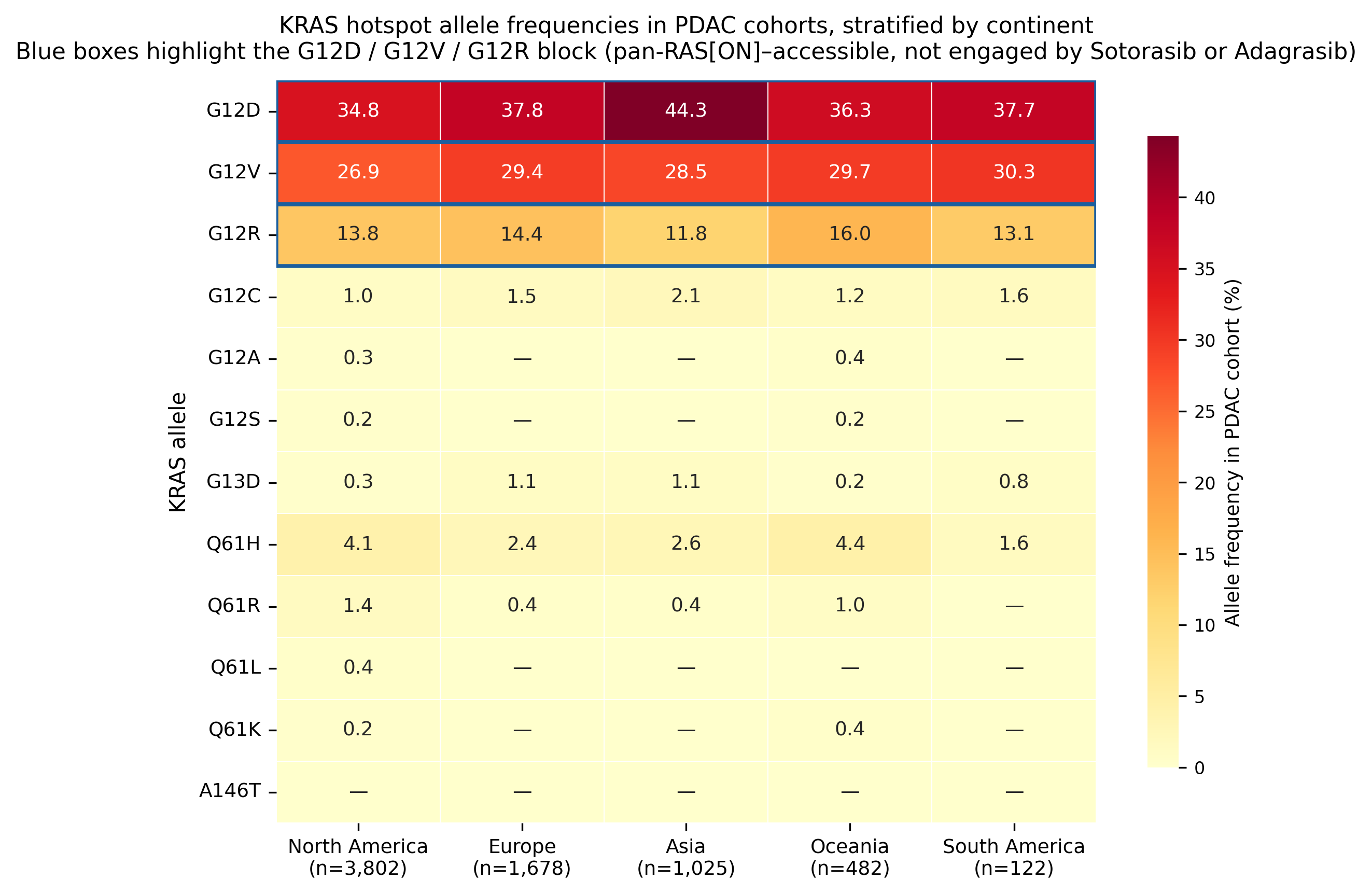
For researchers, that visual pattern is the central fact of the case study. The addressable biology in PDAC is not G12C. It is the G12D, G12V, G12R, and Q61 block.
The stacked distribution makes the same point in a different way. The red G12C segment is a thin slice across continents, while the pan-RAS(ON)-addressable region occupies most of the PDAC bar.
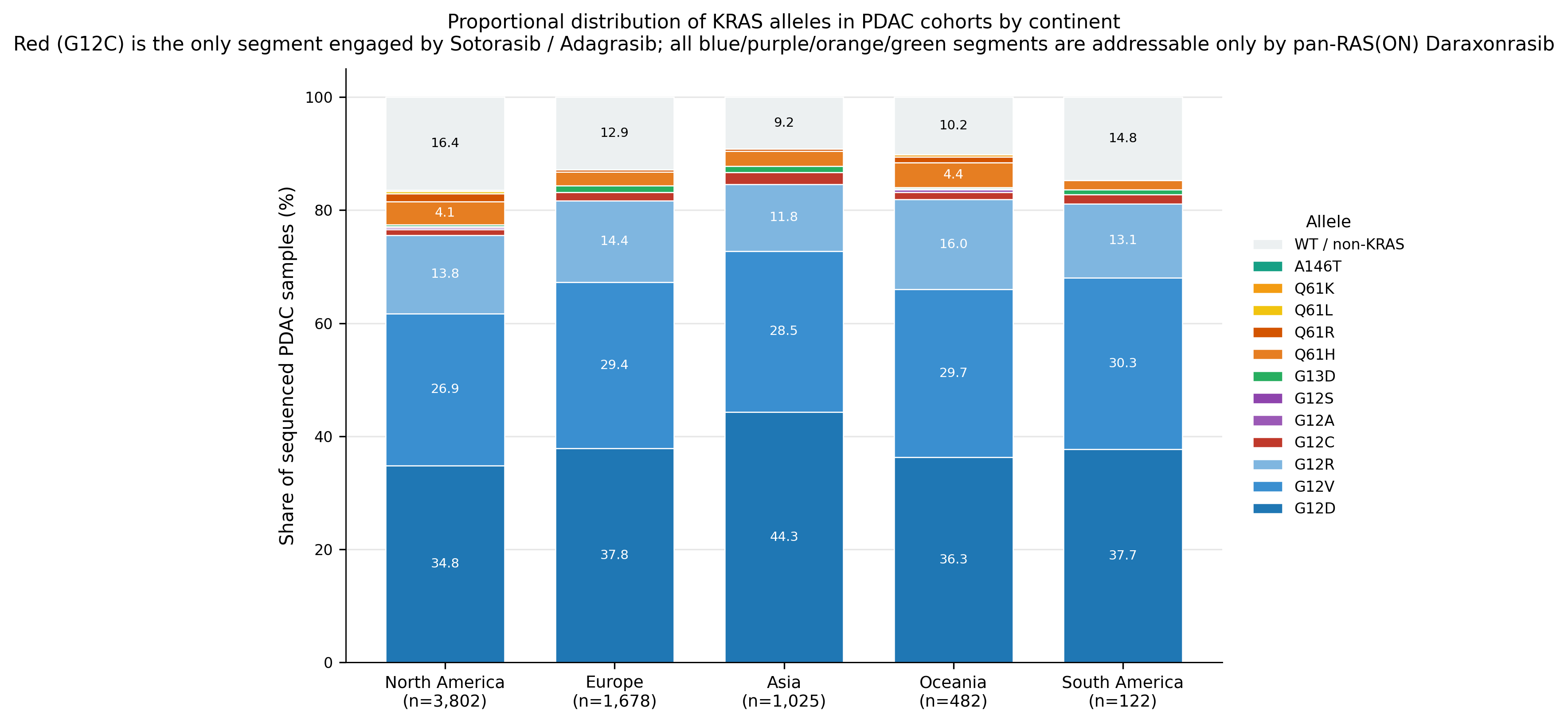
That distinction changes how one should read the RAS drug-development landscape. A G12C inhibitor can be scientifically important and still have limited population reach in PDAC. A broader RAS mechanism may have a much larger translational footprint if its pharmacology is real and its clinical activity holds.
Eligibility: 89.6% Versus 1.4% in Tissue PDAC
K-Dense Web then mapped each observed allele into a coverage class.
The G12C-only class corresponded to the established covalent KRAS-G12C inhibitors. The pan-RAS(ON) class included common G12, G13, and Q61 RAS-family hotspots that would be plausibly engaged by daraxonrasib's active-state mechanism. A third class captured uncovered RAS variants, including atypical A146 variants, remote in-frame indels, and generic "Other" bins. RAS-negative tumors were reported separately.
The headline result was stark:
| Group | Daraxonrasib eligible | G12C inhibitor eligible | Incremental gain |
|---|---|---|---|
| Global tissue PDAC, n = 6,697 | 89.6% | 1.4% | +88.2 percentage points |
| Asia | 91.1% | 2.1% | +89.0 percentage points |
| Europe | 88.1% | 1.5% | +86.6 percentage points |
| North America, tissue | 89.6% | 1.1% | +88.5 percentage points |
| Oceania | 91.7% | 1.2% | +90.5 percentage points |
| South America | 86.1% | 1.6% | +84.4 percentage points |
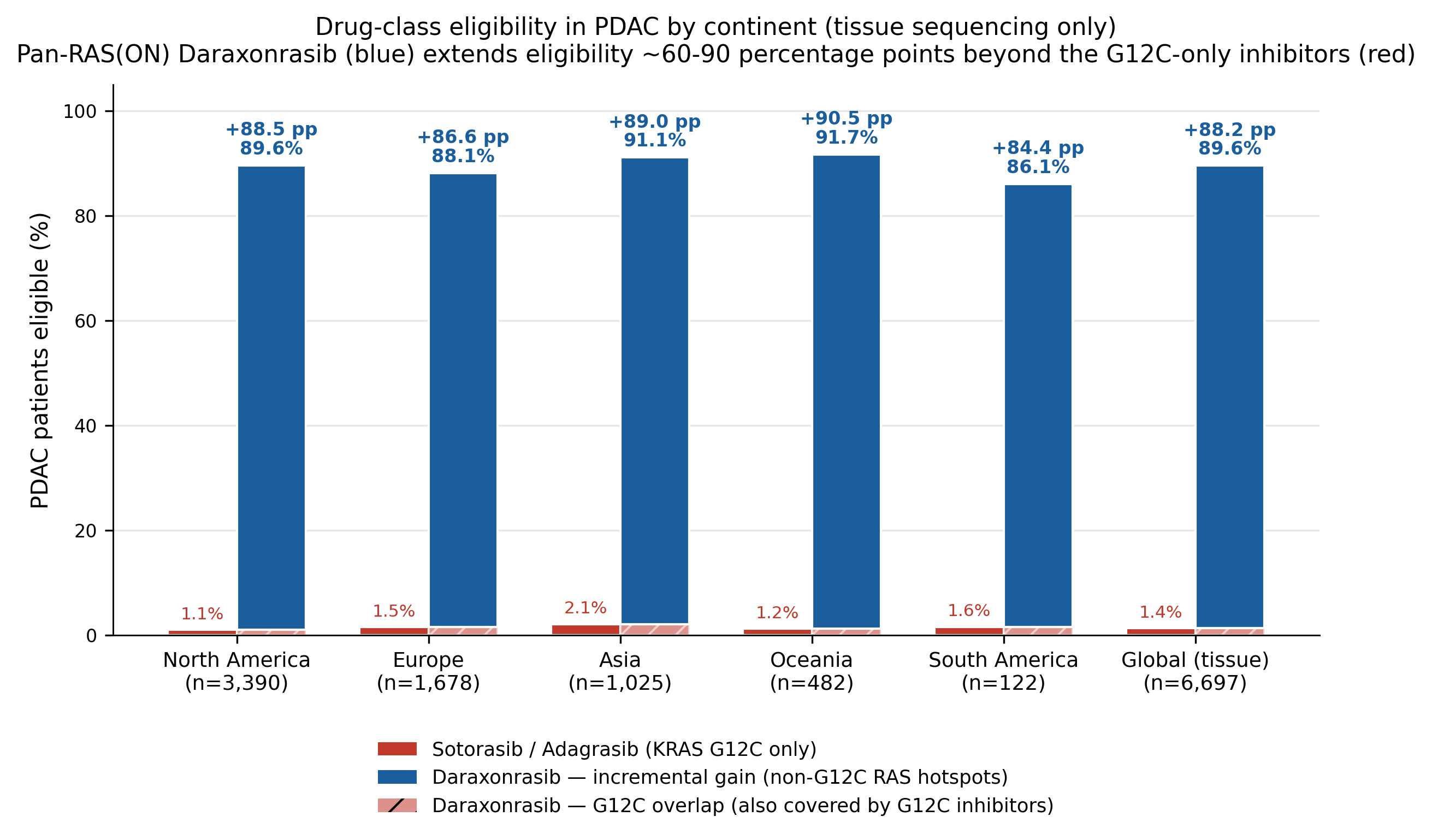
The global tissue estimate implies a roughly 60-90x relative increase in addressable PDAC population for a pan-RAS(ON) strategy compared with G12C-only inhibition. The exact multiplier varies by continent because G12C is rare everywhere and small denominator changes move the ratio, but the qualitative conclusion does not change.
For a translational scientist, this is the value of allele-specific epidemiology. The mechanism, the clinical strategy, and the target population all depend on the same denominator.
Mechanism: OFF-State Covalent Binding Versus ON-State Tri-Complex Pharmacology
K-Dense Web also generated a structured mechanism comparison and rendered it as a schematic. This mattered because eligibility is only meaningful if the binding logic is biologically plausible.
Sotorasib and adagrasib bind KRAS-G12C in the inactive GDP-bound state. They exploit the switch-II pocket and form a covalent bond with the mutant cysteine at codon 12. That chemistry explains both their selectivity and their limitation. Without Cys12, the warhead has no equivalent target.
Daraxonrasib is different. It is a non-covalent pan-RAS(ON) inhibitor built around a cyclophilin A mediated tri-complex. The drug first binds cyclophilin A, and the resulting binary complex docks onto active, GTP-bound RAS at the switch-I/switch-II interface. Because the interaction surface does not require a G12C cysteine, the mechanism can in principle engage common KRAS, NRAS, and HRAS hotspots across codons 12, 13, and 61.
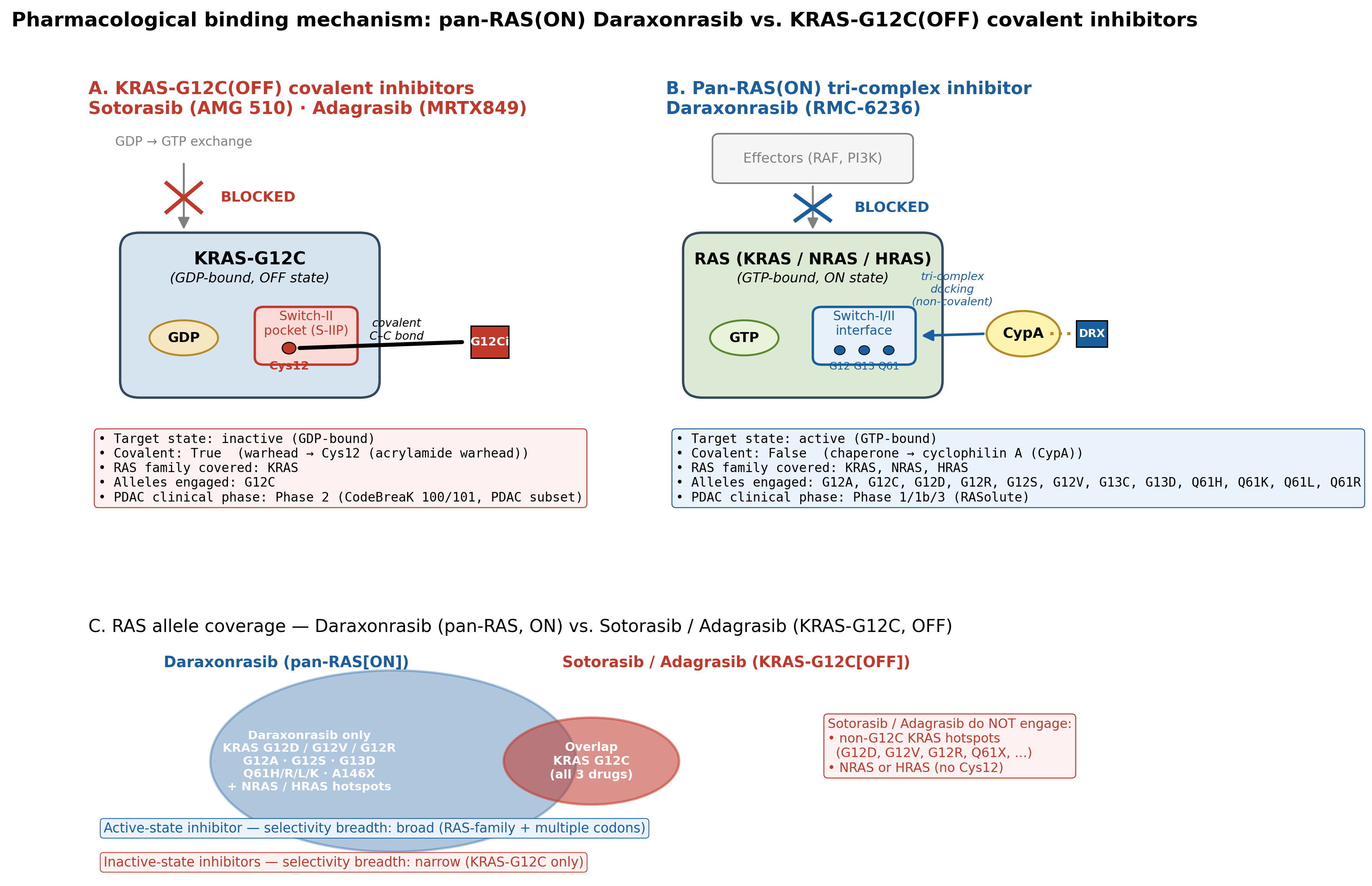
For scientists, this is more than a cartoon. It links chemical mechanism to epidemiology. The G12C drugs are narrow because their chemical handle is narrow. The pan-RAS(ON) strategy is broad because its binding surface is not allele-locked in the same way.
Cross-Continent Statistics: Significant, but Small Effects
The workflow did not stop at descriptive prevalence. K-Dense Web tested whether major allele frequencies differed across continents using the tissue-PDAC cohort.
For each allele, it built a 2 x 5 contingency table: has-allele versus no-allele across five continents. It used Pearson chi-square tests when expected counts were adequate and Fisher-Freeman-Halton exact tests with Monte Carlo simulation when counts were sparse. Pairwise continent comparisons used two-sided Fisher exact tests. Benjamini-Hochberg false-discovery correction was applied across the omnibus family and within each pairwise allele family.
Three alleles showed significant cross-continent distributional differences after correction:
| Allele | Test result | Adjusted significance | Effect size |
|---|---|---|---|
| Q61X | p = 7.6e-10, q = 6.1e-09 | Significant | Cramer's V = 0.085 |
| G12D | p = 4.3e-04, q = 1.7e-03 | Significant | Cramer's V = 0.055 |
| G13D | p = 9.5e-03, q = 2.5e-02 | Significant | Cramer's V = 0.047 |
The interpretation was deliberately conservative. Q61X was enriched in North America compared with Asia and Europe. G12D was highest in Asia, with a roughly 7 percentage point excess over North America and Europe. G13D was rare everywhere but enriched in Europe relative to North America.
However, all effect sizes were small, with Cramer's V below 0.1. That means the differences are statistically detectable in a multi-thousand-sample cohort, but they do not overturn the main clinical conclusion. Across continents, the dominant PDAC population remains pan-RAS(ON)-addressable and largely invisible to G12C-only drugs.
This is a useful example of how K-Dense Web can help scientists avoid two opposite errors: ignoring real heterogeneity, or exaggerating small but significant effects.
Trial Landscape: The Biology Is Reflected in the Pipeline
The final step mapped the RAS-targeted clinical-trial landscape using the ClinicalTrials.gov v2 API. K-Dense Web queried 64 terms across drug classes and synonyms, then filtered and annotated records for pancreatic cancer and advanced solid tumor relevance.
The workflow identified 137 RAS-targeted trials relevant to pancreatic cancer or advanced solid tumors:
| Dimension | Result |
|---|---|
| Total trials | 137 |
| Phase 1 | 58 |
| Phase 1/2 | 47 |
| Phase 2 | 22 |
| Phase 3 | 6 |
| Combination strategies | 107 trials, 78% |
| Monotherapy strategies | 30 trials, 22% |
| Pancreatic cancer only | 34 trials |
| Advanced solid tumors only | 69 trials |
| Both PDAC and solid tumor scope | 32 trials |
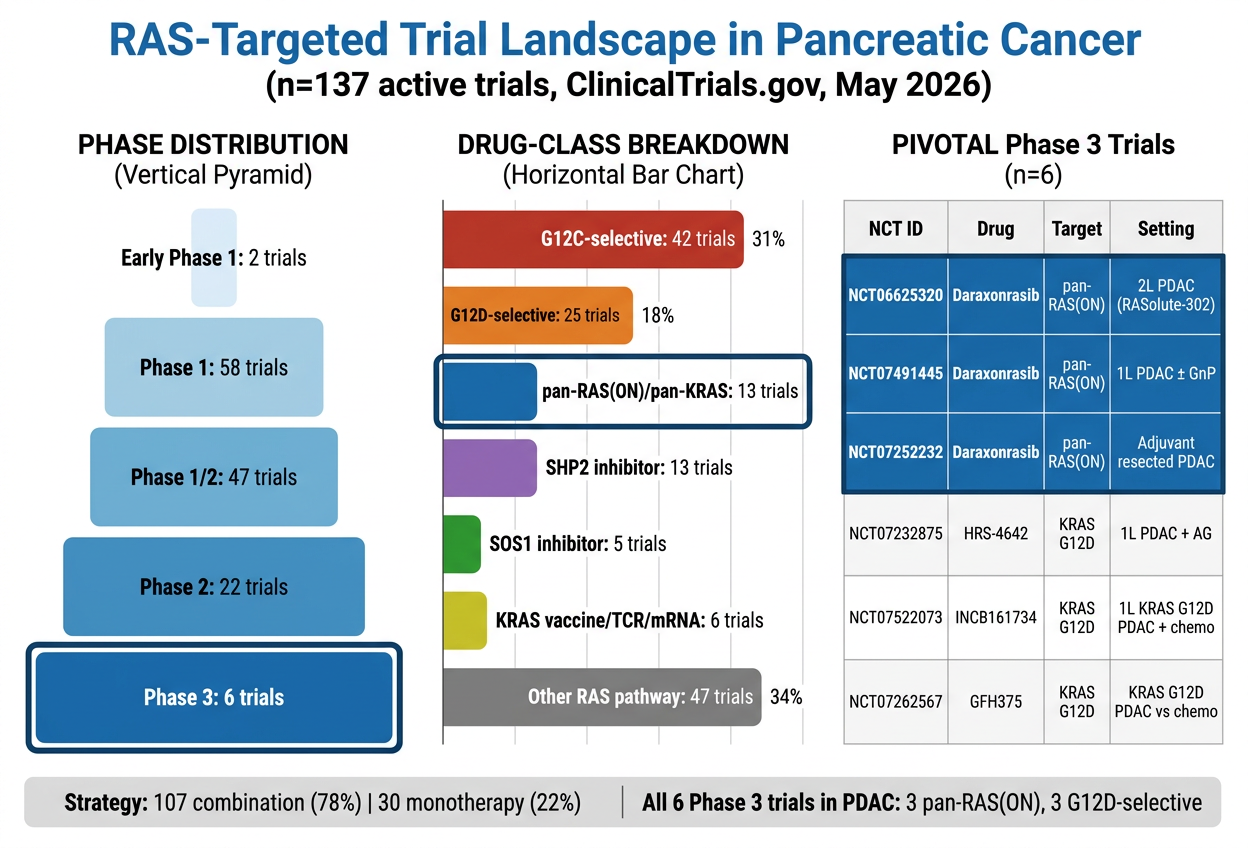
The Phase 3 landscape mirrored the genomic findings. All six Phase 3 trials were registration-relevant pancreatic cancer studies. Three involved daraxonrasib across second-line metastatic, first-line metastatic, and adjuvant settings. The other three were KRAS-G12D-selective programs, reflecting the high prevalence of G12D in PDAC.
That is exactly what one would expect from a rational precision oncology pipeline. G12C has been clinically productive but population-limited in PDAC. G12D and pan-RAS(ON) strategies are where the broader pancreatic cancer opportunity sits.
The trial map also showed that most programs are combination studies. This is biologically unsurprising. RAS signaling is adaptive, feedback-rich, and embedded in a stromal and immune environment that is hostile to single-agent durability. The session identified recurring partner classes, including MEK and ERK inhibitors, SHP2 and SOS1 inhibitors, EGFR antibodies, immune checkpoint inhibitors, and standard chemotherapy backbones such as gemcitabine plus nab-paclitaxel and mFOLFIRINOX.
What Remains Uncovered
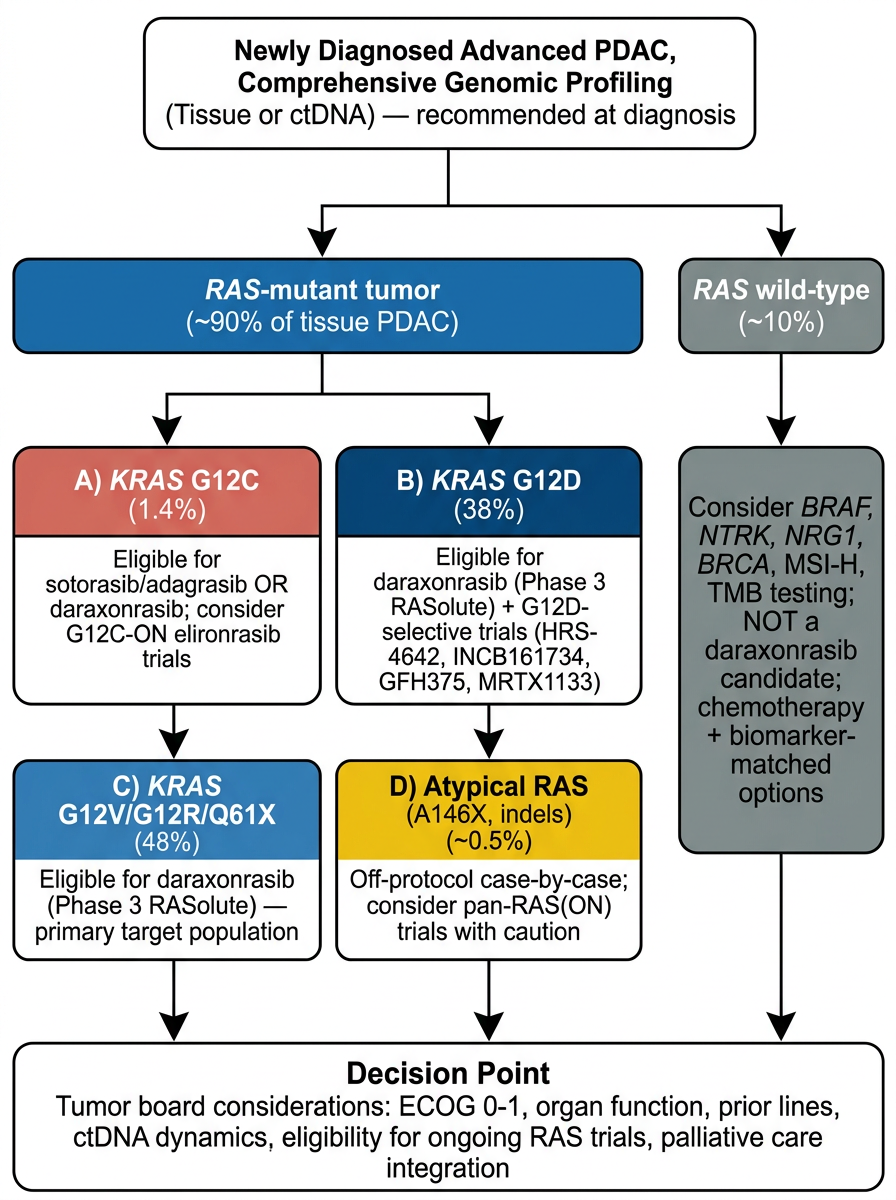
A scientifically useful brief should not only say who is eligible. It should say who is still left out.
K-Dense Web identified several residual gaps:
- RAS-wild-type PDAC. Roughly 10% of tissue PDAC lacks a canonical RAS hotspot. These tumors may be driven by alternative alterations and require separate molecular triage.
- Atypical RAS variants. A146 variants, remote in-frame indels, and poorly annotated "Other" bins may not be covered by current drug classes.
- Liquid-biopsy false negatives. The ctDNA cohort showed much lower apparent eligibility, likely reflecting assay sensitivity and shedding rather than true biology.
- Frail patients. Patients with poor performance status are often excluded from registrational trials, leaving uncertainty about real-world tolerability.
- CNS metastases. Brain penetration and intracranial activity remain separate questions.
- Resistance. Even a broad pan-RAS inhibitor will face evolutionary escape through pathway reactivation, acquired mutations, lineage plasticity, or bypass signaling.
This section is important because it keeps the use case honest. A broad mechanism can change the treatment denominator, but it does not eliminate the need for careful diagnostics, resistance monitoring, and trial enrollment strategy.
Why This Is a K-Dense-Shaped Problem
This session is a good example of the difference between an AI answer and an AI research workflow.
A single model response can summarize that KRAS is common in pancreatic cancer. It can probably mention that G12D is common and G12C is rare. But a scientist or translational team needs more than that. They need:
- Cohort provenance.
- Explicit denominators.
- Allele-level parsing.
- Separate handling of tissue and ctDNA cohorts.
- Geography-aware aggregation.
- Audit tables for drug-coverage assumptions.
- Statistical testing with multiple-comparison correction.
- Mechanism figures grounded in structured data.
- Clinical-trial search and deduplication.
- A written brief that preserves uncertainty.
K-Dense Web handled those as a coordinated research operation. Each step created artifacts the next step could inspect. The eligibility figure depended on the allele table. The mechanism schematic depended on the structured pharmacology JSON. The statistical section depended on tissue-only contingency tables. The final brief depended on the full chain of outputs, not on a disconnected narrative.
For scientists and researchers, that is the central value: the system can keep claim, data, code, figure, and prose connected.
Reproducibility and Review
The session preserved its implementation scripts, logs, intermediate files, and final outputs. The core analysis scripts included:
01_fetch_genomic_data.pyfor cBioPortal and literature-cohort aggregation.03_gap_analysis_eligibility.pyfor drug-class eligibility.04_generate_visualizations.pyfor publication figures.05_statistical_comparison.pyfor cross-continent tests.06_clinical_trial_landscape.pyfor ClinicalTrials.gov acquisition and annotation.
The final deliverable was a 19-page tumor-board technical brief with 39 verified references, eight figures, and a peer-review report. The internal review accepted the brief with minor revisions and highlighted the strongest parts of the workflow: mechanism clarity, quantitative eligibility math, continent stratification, trial-landscape completeness, and a candid "left out" section.
The limitations were also explicit. South America was represented by a single small cohort. Some rare-allele enrichments may reflect sequencing-panel or cohort artifacts. Phase 3 daraxonrasib survival data were treated as press-release based pending full peer-reviewed publication. A journal-submission version would benefit from deeper systematic-review formalism and pharmacoeconomic analysis.
That is how scientific AI output should look. Not omniscient, not frictionless, and not detached from uncertainty. Reproducible, inspectable, and clear about the assumptions that matter.
What Researchers Can Take Away
The biological takeaway is straightforward: in PDAC, the clinically meaningful RAS population is much larger than KRAS-G12C. In this session's tissue-based global analysis, pan-RAS(ON)-addressable alleles covered 89.6% of PDAC samples, while G12C-only inhibitors covered 1.4%. That difference follows directly from the allele distribution: G12D, G12V, and G12R dominate, while G12C is consistently rare.
The methodological takeaway is broader. Scientific research increasingly requires workflows that can move across APIs, literature, statistics, figures, and final communication without losing provenance. K-Dense Web is useful when the question has many linked parts and when the answer needs to be defensible, not just fluent.
For a cancer biologist, this session offers an allele-level map of RAS biology in PDAC. For a translational scientist, it connects that map to therapeutic mechanism and patient eligibility. For a clinical-trial strategist, it shows why pan-RAS(ON) and G12D-selective programs dominate the late-stage PDAC pipeline. For a researcher evaluating agentic AI, it shows what a complete computational research session can produce when the system is asked to do more than summarize.
Download the full technical brief
Generated using K-Dense Web (k-dense.ai). This post summarizes a research workflow and is not medical advice.
