Most scientific Python code runs on the CPU. NumPy, pandas, scikit-learn, SciPy, NetworkX: the standard stack is CPU-only by default. Meanwhile, NVIDIA GPUs capable of processing thousands of operations in parallel sit idle because the software bridge between a scientist's Python script and the GPU hardware is genuinely hard to cross.
The gap isn't about intelligence or effort. It's about specialization. Writing GPU-optimized code requires understanding CUDA programming, GPU memory hierarchies, asynchronous execution, kernel launch overhead, host-device data transfer costs, and a dozen library-specific APIs, each with their own installation quirks, conventions, and pitfalls. That's a full engineering discipline, separate from the scientific work itself.
We built the optimize-for-gpu skill so scientists can get GPU-level performance without becoming GPU engineers.
The NVIDIA Ecosystem: Powerful, but Complex
NVIDIA's GPU-accelerated Python ecosystem is remarkably comprehensive. Twelve libraries cover nearly every scientific computing domain:
| Library | Replaces | Domain |
|---|---|---|
| CuPy | NumPy, SciPy | Array math, linear algebra, FFT, signal processing |
| Numba CUDA | Custom loops | Custom GPU kernels, fine-grained thread control |
| Warp | Simulation loops | Physics simulation, mesh operations, differentiable programming |
| cuDF | pandas | DataFrame operations, ETL, groupby, joins |
| cuML | scikit-learn | Classification, regression, clustering, dimensionality reduction |
| cuGraph | NetworkX | PageRank, centrality, community detection, shortest paths |
| cuCIM | scikit-image | Image filtering, morphology, segmentation, digital pathology |
| cuVS | Faiss, Annoy | Vector search, nearest neighbors, RAG retrieval |
| cuSpatial | GeoPandas | Spatial joins, distance calculations, trajectory analysis |
| KvikIO | numpy.fromfile | GPUDirect Storage, S3/HTTP to GPU, binary file IO |
| cuxfilter | matplotlib | Interactive cross-filtering dashboards on GPU |
| RAFT | scipy.sparse.linalg | Sparse eigensolvers, device memory, multi-GPU primitives |
That's twelve libraries. Some, like CuPy and the RAPIDS suite (cuDF, cuML, cuGraph), dispatch to NVIDIA's hand-tuned CUDA libraries (cuBLAS, cuFFT, cuSOLVER, cuSPARSE). Others, like Numba CUDA and Warp, JIT-compile your Python code directly into custom CUDA kernels. Either way, the performance is there. The problem is everything else: knowing which library covers which operation, how to install it, how to manage GPU memory allocation, when to synchronize the device, how to minimize expensive CPU-GPU data transfers, how to handle operations that have no GPU equivalent, and how to compose multiple libraries together without redundant copies.
A researcher who wants to accelerate a correlation matrix computation shouldn't need to know that CuPy dispatches to cuBLAS under the hood, that GPU operations are asynchronous and require explicit synchronization for accurate timing, or that transferring a large result back to CPU for a subsequent step that lacks a GPU implementation needs to happen at exactly the right point in the pipeline. They should describe what they need and get working, optimized code.
What optimize-for-gpu Does
The optimize-for-gpu skill is an Agent Skill that functions as a GPU optimization engineer embedded in your AI coding assistant. Give it your existing CPU-bound Python code, or describe a computation you want to build from scratch, and it handles the GPU engineering.
The skill carries detailed knowledge of all twelve libraries: their APIs, their performance characteristics, their interoperability patterns, and their failure modes. When it encounters your code, it follows a systematic process:
-
Assess the workload. It identifies what your code actually does (array math, dataframe operations, ML training, graph analytics, image processing, physics simulation, vector search, geospatial analysis, file IO) and determines which parts are compute-bound versus IO-bound.
-
Select the right tools. Based on the workload, it picks the optimal GPU library or combination of libraries. It knows that a pandas groupby maps to cuDF, a scikit-learn pipeline maps to cuML, a particle simulation maps to Warp, and a custom algorithm with complex per-element logic needs a Numba CUDA kernel. More importantly, it knows when a workload spans multiple libraries and how to compose them without unnecessary data copies.
-
Write the GPU code. It produces GPU-accelerated code that handles memory management, device synchronization, data transfer, warm-up, and GPU-specific optimizations like pre-allocating output arrays, batching small operations to amortize kernel launch overhead, and using float32 where precision allows for higher throughput.
-
Handle the hard cases. Real scientific code rarely maps cleanly to a single library. When part of a pipeline has no GPU equivalent (like hierarchical clustering or connected-component labeling), the skill implements hybrid strategies, accelerating the expensive bottleneck on GPU and managing the transfer to CPU for the remainder. It also restructures serial Python loops into batched GPU operations, replacing per-element iteration with vectorized computation across the entire dataset simultaneously.
The result is that a scientist describes a problem or provides existing code, and gets back optimized GPU code that would have taken a CUDA engineer hours or days to write. The skill absorbs the complexity of the NVIDIA ecosystem so the scientist doesn't have to.
Benchmarks: 15 Workloads, 58x Average Speedup
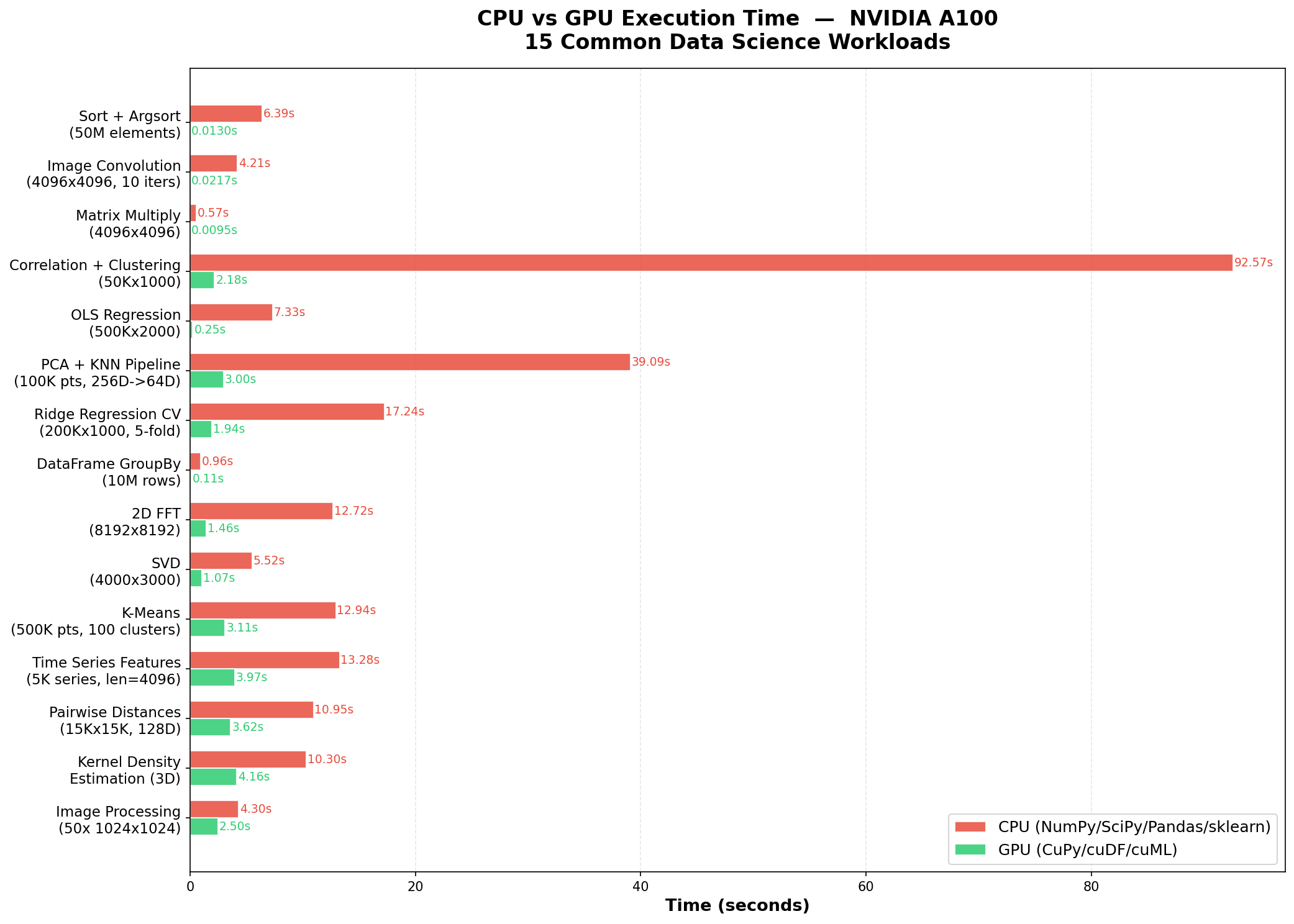
To measure how well this works in practice, we benchmarked the skill on 15 common data science workloads. We wrote 15 standard CPU scripts using NumPy, SciPy, pandas, and scikit-learn, then used the optimize-for-gpu skill to produce GPU equivalents using CuPy, cuDF, and cuML. Both versions ran on identical hardware via Modal cloud infrastructure.
Hardware: NVIDIA A100 (40 GB HBM2e), 4 CPU cores, 16 GB RAM. Each script reports core computation time only, excluding data generation and library imports. GPU scripts include a warm-up pass to exclude one-time CUDA initialization costs.
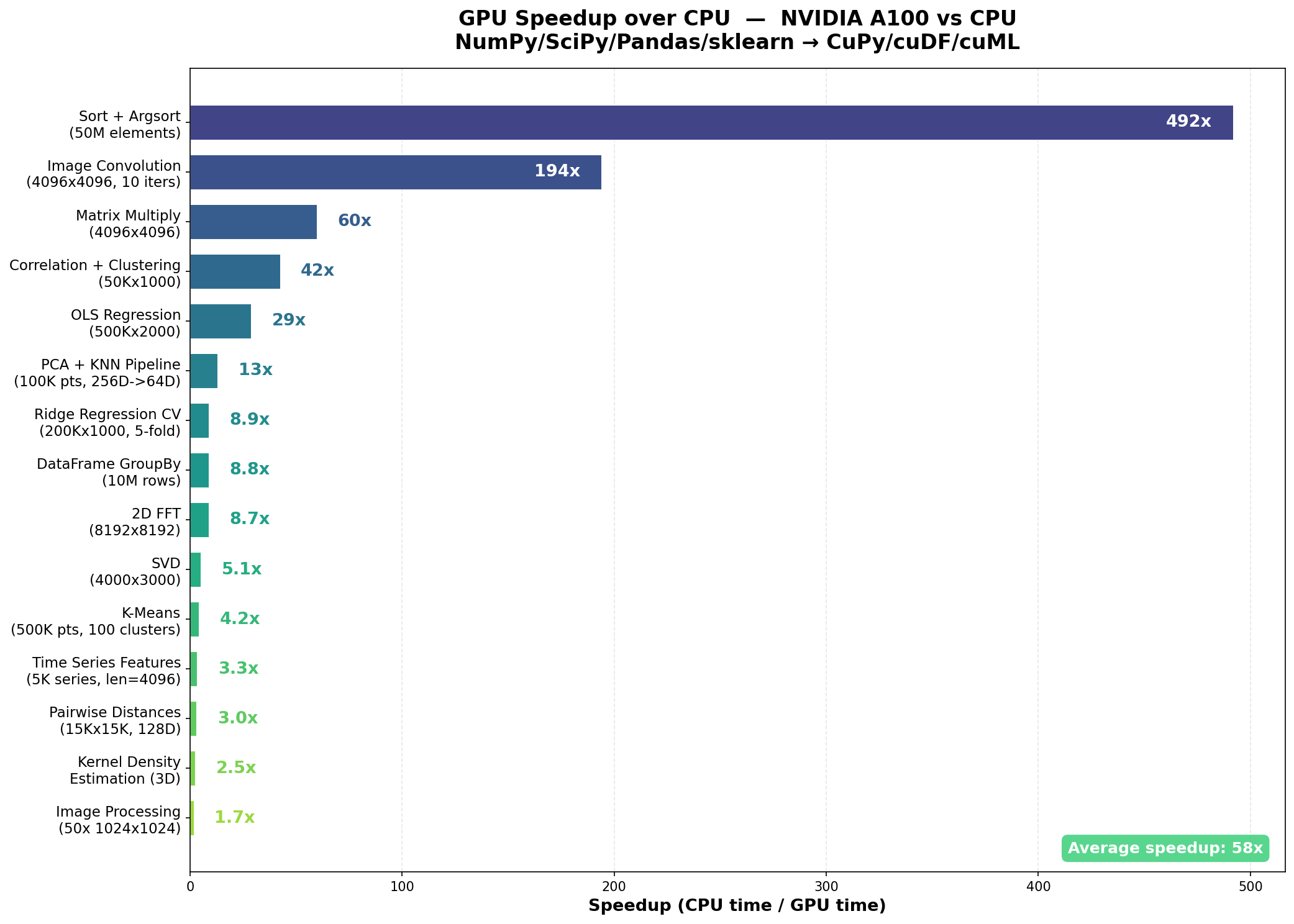
Average speedup across all 15 benchmarks: 58x, ranging from 1.7x to 492x.
The standout results:
- Sort + Argsort (50M elements): 492x, from 6.39s to 0.013s
- Image Convolution (4096x4096, 10 iterations): 194x, from 4.21s to 0.022s
- Matrix Multiply (4096x4096): 60x, from 0.57s to 0.010s
- Correlation + Hierarchical Clustering (50Kx1000): 42x, from 92.57s to 2.18s
- OLS Regression (500Kx2000): 29x, from 7.33s to 0.25s
The multi-library pipeline benchmarks are particularly telling. These chain multiple operations together, reflecting how data scientists actually work: not isolated operations, but multi-step workflows where data flows from one computation to the next.
- PCA + KNN Pipeline (100K points, 256D to 64D): 13x
- Ridge Regression with 5-fold CV (200Kx1000): 8.9x
- Time Series Feature Extraction (5K series): 3.3x
Why Speedups Vary
The range from 1.7x to 492x reflects the diversity of these workloads and the real engineering challenges in GPU optimization.
Highest speedups (Sort, Convolution, MatMul, Correlation): These are highly parallelizable operations with regular memory access patterns. The A100's 1,555 GB/s memory bandwidth and thousands of CUDA cores can process massive arrays orders of magnitude faster than a CPU. The correlation + clustering benchmark saw 42x because the bottleneck, a 50K x 1000 correlation matrix, maps to a single GPU matrix multiply.
Moderate speedups (OLS, PCA+KNN, Ridge CV, GroupBy, FFT): These involve a mix of compute-bound and memory-bound phases. GPU acceleration helps significantly, but some steps (iterative alpha sweeps, data indexing for cross-validation folds, hash-based grouping) are harder to parallelize.
Lower speedups (K-Means, Time Series, Pairwise Distances, KDE, Image Pipeline): These involve iterative algorithms (K-Means convergence), memory-constrained batching (KDE), or operations already well-optimized on CPU. The image processing pipeline had the lowest speedup (1.7x) because connected-component labeling is inherently sequential and required CPU fallback.
The skill handled all of these cases, including the hard ones. For the correlation + clustering benchmark, it recognized that hierarchical clustering has no GPU equivalent, so it computed the correlation matrix on GPU (the bottleneck), then transferred the result to CPU for the clustering step. For time series feature extraction, it replaced a per-series Python loop over 5,000 time series with fully vectorized batch FFT, cumulative-sum-based rolling statistics, and batch autocorrelation across all series simultaneously on GPU. These are the kinds of decisions that require deep knowledge of which operations are GPU-friendly and which aren't, and that is exactly what the skill provides.
Beyond the Benchmarks
The 15 benchmarks cover a representative slice of data science, but the skill's scope extends across the full NVIDIA ecosystem:
- Graph analytics: PageRank, community detection, betweenness centrality on networks with millions of edges via cuGraph
- Vector search: Approximate nearest neighbor search for RAG pipelines, recommender systems, and embedding retrieval via cuVS
- Medical imaging: Whole-slide image processing, cell segmentation, H&E stain normalization via cuCIM
- Geospatial analysis: Point-in-polygon tests, spatial joins on millions of GPS coordinates, trajectory reconstruction via cuSpatial
- Physics simulation: Particle systems, fluid dynamics, cloth simulation, differentiable rendering via Warp
- Interactive dashboards: Cross-filtering visualization on million-row datasets via cuxfilter
- High-performance IO: Loading binary data directly from disk or S3 into GPU memory, bypassing CPU entirely, via KvikIO
- Sparse eigensolvers: Spectral methods and graph partitioning on large sparse matrices via RAFT
All twelve libraries interoperate through the CUDA Array Interface, which allows zero-copy data sharing between CuPy, cuDF, cuML, cuGraph, PyTorch, JAX, and the rest. The skill knows these integration patterns and uses them to build end-to-end pipelines that keep data on the GPU throughout, avoiding the costly round-trips through CPU memory that can erase the performance gains.
Getting Started
The optimize-for-gpu skill is available through three channels:
Scientific Agent Skills (free, open source). Compatible with any AI agent that supports Agent Skills, including ChatGPT, Claude.ai, Claude CoWork, Claude Code, Codex, Gemini CLI, and Cursor. Install the skills, mention GPU acceleration or describe a compute-intensive workload, and the skill activates automatically. Browse the repository on GitHub.
K-Dense BYOK (free, open source). Our desktop AI co-scientist includes optimize-for-gpu alongside 170+ other scientific skills. Bring your own API keys, choose from 40+ models, and run everything locally. Get started on GitHub.
K-Dense Web (full platform). Cloud GPUs, persistent sessions, and end-to-end research pipelines. Upload your code, describe the optimization goal, and the platform handles execution on cloud GPU infrastructure. Try it at www.k-dense.ai.
Your Code, Faster
If you're running numerical Python code on a CPU and you have access to an NVIDIA GPU, whether on your workstation, in the cloud, or through a platform like Modal, there's a good chance the optimize-for-gpu skill can deliver a meaningful speedup. Sometimes it's 2x. Sometimes it's 492x. The skill figures out which libraries to use, writes the GPU code, handles the memory management and synchronization, and manages the hybrid CPU-GPU boundary when needed.
The hard part of GPU programming isn't the concept. It's the hundreds of practical decisions about which library to use, how to manage device memory, when to synchronize, and how to compose operations efficiently. The optimize-for-gpu skill makes those decisions for you, so you can focus on the science.
Questions? Reach out at contact@k-dense.ai.
Related Resources:
