Hantavirus research is not a single-lane problem. It sits at the intersection of rodent ecology, climate, rural housing, occupational exposure, clinical recognition, surveillance systems, and public health communication. That is exactly the kind of problem K-Dense Web was built to work on.
The basic story is simple enough to explain in one sentence: hantaviruses are carried by rodents, people are usually infected through contact with contaminated urine, droppings, or saliva, and severe disease can appear after exposure CDC. The research story is harder. Which rodents matter in which places? Which weather patterns change risk? Which case counts are final and which are provisional? Which prevention recommendations are sturdy enough for public messaging? Which open questions are evidence gaps rather than speculation?
That is where an agentic research workflow becomes useful. Instead of asking one model for a summary, a researcher can ask K-Dense Web to build an evidence map, check it against primary sources, generate figures, flag uncertainty, and produce publishable briefings for different audiences: scientists, clinicians, policy teams, health departments, journalists, educators, and the public.
Why hantavirus is a K-Dense-shaped problem
Hantavirus is rare enough in many countries that expertise is unevenly distributed, but serious enough that delayed recognition matters. CDC notes that hantavirus pulmonary syndrome can begin with fever, fatigue, and muscle aches, then progress four to ten days later to coughing and shortness of breath as the lungs fill with fluid CDC. CDC also states that 38% of people who develop respiratory symptoms may die from the disease CDC.
The global picture is broader than one syndrome. WHO describes hantaviruses as a group of rodent-borne viruses that can cause hemorrhagic fever with renal syndrome and hantavirus pulmonary syndrome, with risk shaped by human contact with infected rodents and contaminated environments WHO. WHO also emphasizes surveillance, laboratory capacity, risk communication, community engagement, early detection, patient care, outbreak response, and One Health approaches that connect human health, rodent reservoirs, and the environment WHO.
That combination creates a research bottleneck. The evidence lives in different places: CDC clinical pages, NNDSS reporting tables, WHO guidance, ECDC surveillance reports, ecology papers, veterinary and wildlife surveillance, local health department advisories, and older outbreak investigations. Human experts can synthesize it, but the work is slow, repetitive, and easy to under-scope.
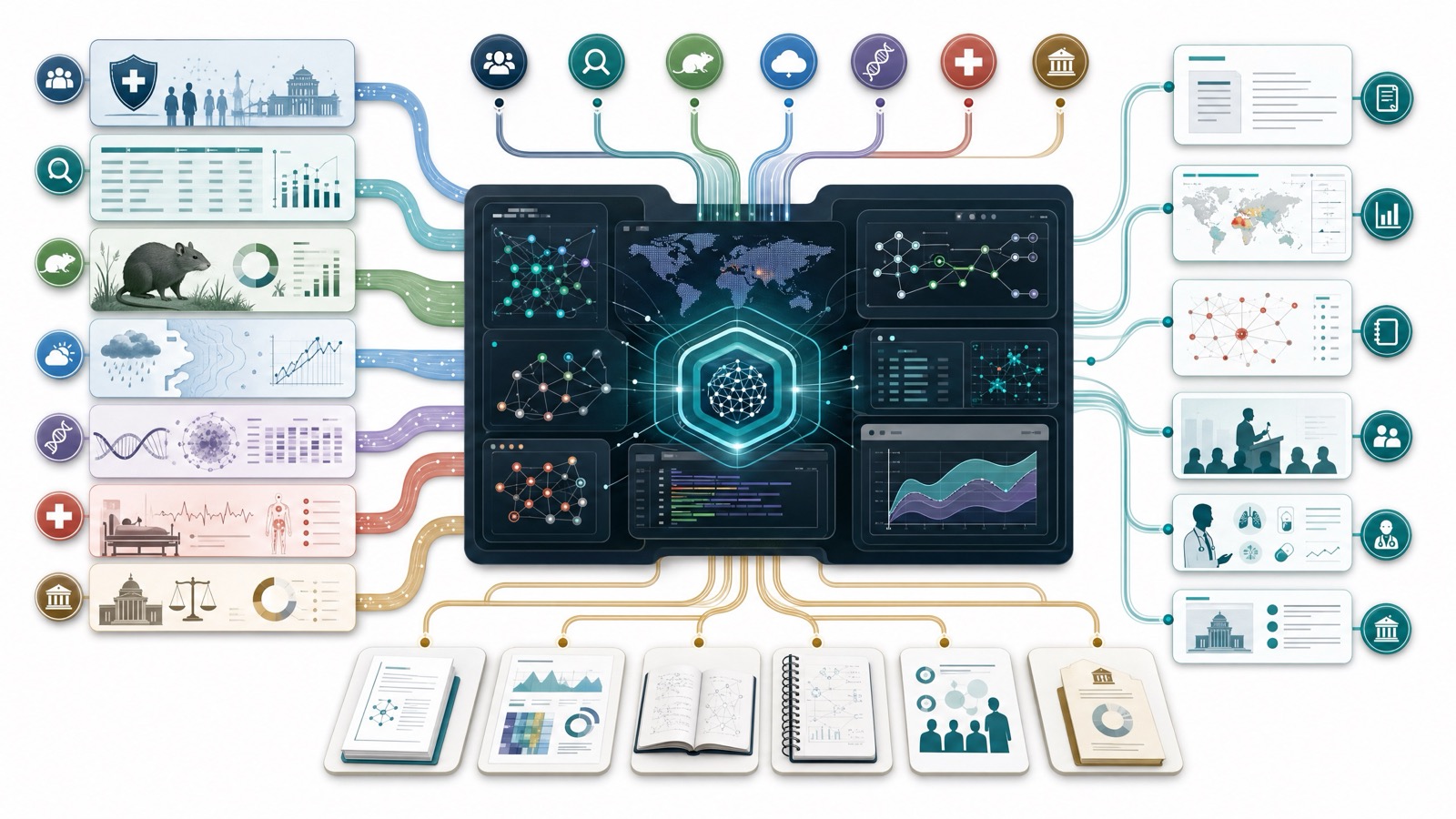
K-Dense Web can make the workflow more like a research operation than a literature search.
The cruise-ship cluster shows why this matters now
The reason this topic feels urgent is the recent cruise-ship cluster. On May 4, 2026, WHO reported a hantavirus cluster linked to cruise ship travel after a severe respiratory illness cluster was reported aboard a ship carrying 147 passengers and crew WHO Disease Outbreak News. As of that WHO update, seven cases had been identified, including two laboratory-confirmed hantavirus cases, five suspected cases, three deaths, one critically ill patient, and three people with mild symptoms WHO Disease Outbreak News.
By May 7, WHO said eight cases had been reported, including three deaths, and five of the eight had been confirmed as hantavirus WHO. WHO also identified the virus involved as Andes virus, the hantavirus species known to be capable of limited human-to-human transmission linked to close and prolonged contact WHO. That does not mean hantavirus behaves like a typical respiratory pandemic virus. WHO assessed the public health risk as low, while noting that more cases could be reported because of the incubation period WHO.
This is exactly the kind of incident where a K-Dense Web workflow is useful. The facts are evolving, the transmission question is nuanced, the response spans multiple countries, and the public needs calm, clear communication. WHO described coordinated international response under the International Health Regulations, shipment of 2,500 diagnostic kits to laboratories in five countries, an expert deployed on board, and operational guidance for safe disembarkation and onward travel WHO. ECDC similarly characterized the event as rapidly evolving and preliminary, with recommendations expected to update as information becomes available ECDC.
For an event like this, K-Dense Web could maintain a living evidence brief: track changing case counts, distinguish confirmed from suspected cases, summarize WHO and ECDC risk assessments, compare media reports against primary public health sources, organize diagnostic and sequencing updates, and generate separate outputs for scientists, travel operators, policy teams, clinicians, and the public.
Beyond synthesis: computational work around the virus
The first use case for K-Dense Web is evidence synthesis, but hantavirus research also needs computation. A useful agent should not stop at reading papers. It should be able to run code, inspect datasets, build models, test assumptions, and turn exploratory analysis into reproducible artifacts.
For virologists and computational biologists, that could mean assembling a pipeline to compare hantavirus genomes across strains, align segments, annotate coding regions, summarize mutations, and generate phylogenetic trees. For a team tracking Sin Nombre virus, Andes virus, Seoul virus, or Puumala virus, K-Dense Web could help organize public sequences, check metadata quality, build reproducible notebooks, and produce figures that show how isolates cluster by geography, host, or collection year.
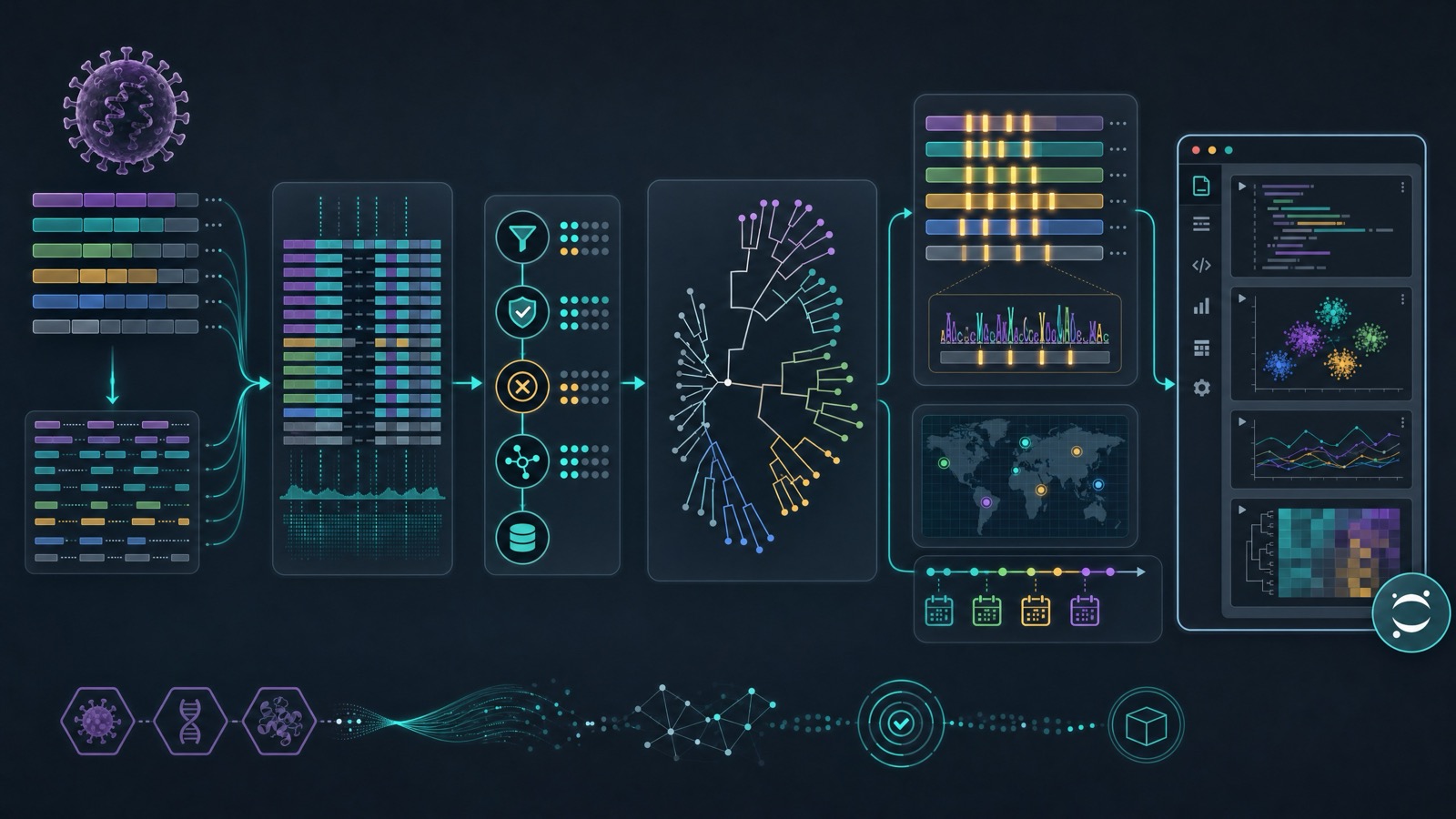

For structural and molecular work, K-Dense Web can coordinate computational tasks around viral proteins: collecting reference sequences, preparing multiple sequence alignments, mapping conserved regions, identifying antigenic or functional motifs, and comparing candidate targets across hantavirus species. That does not replace experimental virology, but it gives wet-lab teams a faster way to frame hypotheses before deciding what to clone, express, test, or prioritize.
For ecology and epidemiology teams, the computational work looks different. K-Dense Web can help merge surveillance records, rodent trapping data, land-use layers, climate variables, and human exposure context into analysis-ready datasets. It can run spatial models, create risk maps, compare model specifications, evaluate lagged climate features, and generate uncertainty-aware visualizations. A 2025 spatial risk study, for example, found that rodent richness, aridity, higher temperatures, and open developed areas were associated with hantavirus risk in U.S. models Wiley. A K-Dense workflow could reproduce the broad analysis pattern on new regions, new data, or updated surveillance windows while keeping the model assumptions visible.
For clinical and public health researchers, K-Dense Web can support computational triage of messy operational data: extracting exposure histories from case reports, standardizing timelines from symptom onset to hospitalization, comparing diagnostic criteria, and building dashboards that separate confirmed cases, suspected cases, and provisional reporting. CDC notes that diagnosing hantavirus infection early can be difficult because initial symptoms can resemble influenza and early testing may need to be repeated after symptom onset CDC. That kind of clinical ambiguity is exactly where structured data extraction and reproducible analysis can help.
The point is not that one agent should be trusted to make final scientific claims alone. The point is that K-Dense Web can turn a broad question into a computational workspace: data ingestion, cleaning, modeling, visualization, literature grounding, and peer review in the same loop.
The prompt
Imagine giving K-Dense Web a prompt like this:
Build a research briefing and computational analysis workspace on hantavirus risk for a scientifically curious public health audience. Cover transmission, clinical course, U.S. and global surveillance, rodent ecology, climate and land-use drivers, prevention, viral genomics, spatial risk modeling, and open research questions. Separate established evidence from hypotheses. Produce a source library, reproducible notebooks, figures, an evidence memo, and a peer-review checklist.
That prompt is deliberately broad. It asks for synthesis, not a single answer. A good workflow would break it into parallel tracks:
- Literature and guideline retrieval from CDC, WHO, ECDC, PubMed, and public health agencies.
- Surveillance extraction from notifiable disease sources, with explicit flags for provisional data.
- Ecology and climate synthesis across rodent reservoirs, land use, precipitation, aridity, and human exposure.
- Computational analysis across sequences, surveillance data, spatial covariates, and clinical timelines.
- Clinical summarization for diagnosis, care pathways, and prevention guidance.
- Output generation, including figures, notebooks, citations, and peer review.
The value is not that the agent "knows" hantavirus. The value is that it can keep many moving parts alive at once, preserve provenance, and turn intermediate research into reusable artifacts.
From analysis to communication, science, and policy
The right output is not a one-off presentation. It is a research package that a scientist, public health analyst, policy advisor, or communications team can inspect, revise, and reuse. The hard part of scientific communication is not only writing. It is maintaining a chain from claim to source to figure to final narrative, then adapting that narrative for the people who need to act on it.
A strong Hantavirus workflow should produce:
- A scientific evidence memo with claims, citations, uncertainty levels, and open questions.
- A policy briefing for health agencies that summarizes risk, surveillance gaps, prevention priorities, and resource needs.
- A public communication package that explains exposure routes, symptoms, and prevention without sensationalism.
- A clinician-facing brief that emphasizes early recognition, exposure history, diagnostic limitations, and escalation pathways.
- A field-team or occupational-health guide for cabins, farms, parks, construction sites, and rodent-disturbed environments.
- A global map distinguishing hantavirus pulmonary syndrome and hemorrhagic fever with renal syndrome.
- A transmission diagram from rodent reservoir to contaminated dust to human exposure.
- A clinical timeline showing prodrome, cardiopulmonary progression, and recovery.
- A surveillance chart that distinguishes final data from provisional data, since CDC publishes weekly and annual NNDSS hantavirus data and notes that reporting is part of the notifiable disease system CDC.
- Reproducible notebooks for cleaning surveillance data, plotting case trends, and documenting caveats.
- Sequence-analysis workflows for comparing strains, building phylogenies, and identifying conserved viral regions.
- Spatial modeling scripts that combine rodent ecology, land use, climate, and exposure variables.
- A risk matrix for cabins, farms, sheds, fieldwork, and disturbed habitats.
- A prevention graphic based on ventilation, wet cleaning, disinfection, and rodent exclusion guidance CDC.
- A "surprises and open questions" section that separates rare Andes virus person-to-person transmission from the much more common rodent-exposure pathway WHO.
That audience shift matters. A scientific memo can talk about host richness, spillover dynamics, and model uncertainty. A county health advisory needs plain instructions. A policy brief needs tradeoffs: surveillance capacity, lab readiness, public messaging, occupational guidance, and funding priorities. A classroom explainer needs the story without the jargon. K-Dense Web can help generate each version from the same underlying evidence base, so the message changes but the source trail does not.
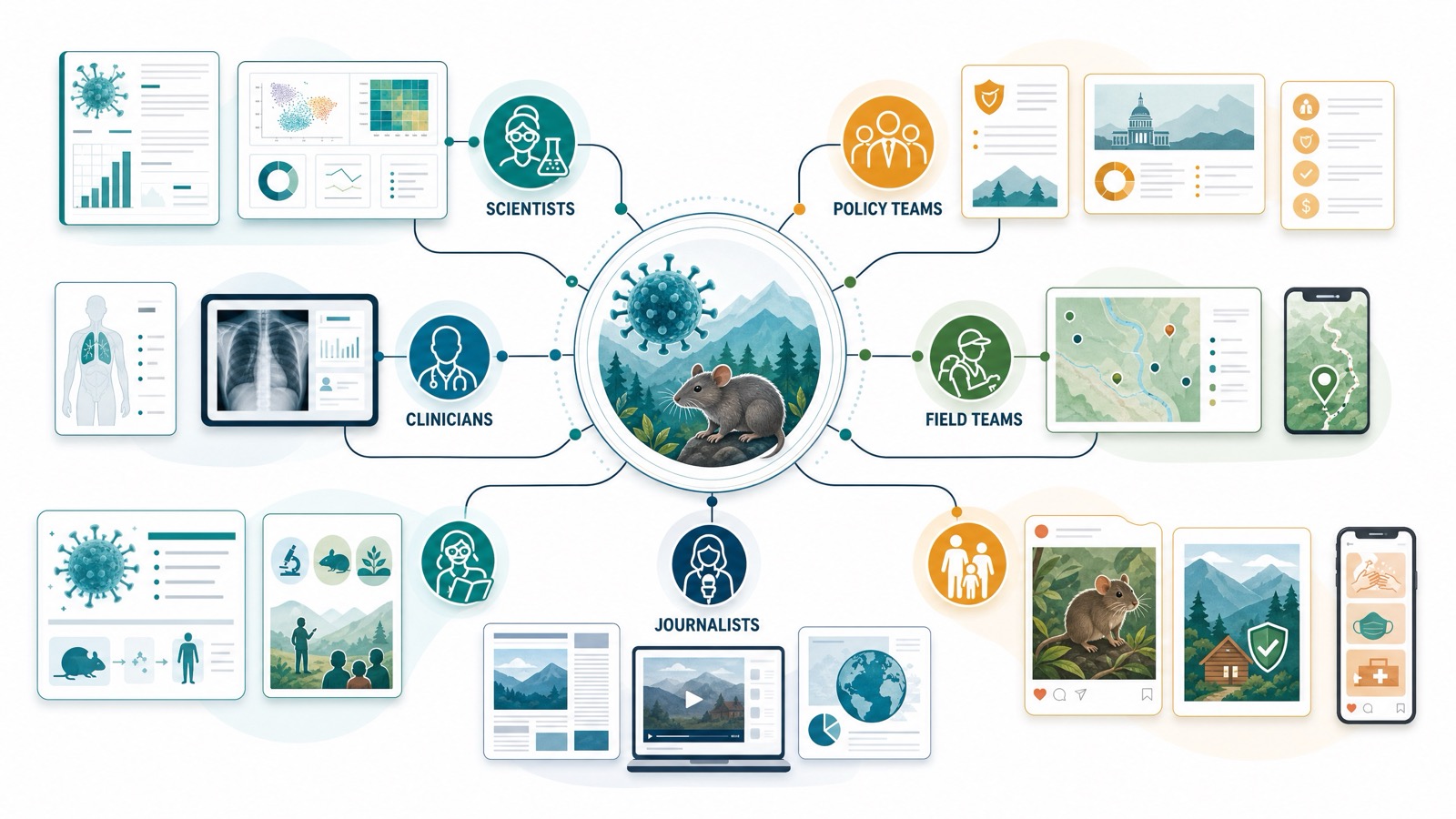
Hantavirus is an ideal test of whether an AI research system can be interesting without becoming sensational. The goal is to make the science memorable while preserving uncertainty.
The research questions K-Dense Web can help organize
The current literature points to several areas where synthesis is genuinely useful.
First, rodent ecology is more complex than a one-host story. A 2025 spatial risk study found that rodent richness was positively associated with hantavirus risk in U.S. models and highlighted recent rodent surveillance showing Sin Nombre virus detections across multiple rodent species in eastern New Mexico Wiley. The same study argues that community-level host dynamics deserve more attention when modeling risk Wiley.
Second, climate and land use matter, but not in a simplistic way. The 2025 study found that low precipitation and higher temperatures were important for structuring spatial hantavirus risk, and its title highlights open developed areas and arid climates as risk-associated features in the western United States Wiley. That kind of result invites a careful K-Dense workflow: extract the model assumptions, compare them with older outbreak ecology, map what is known region by region, and avoid turning correlation into overconfident prediction.
Third, surveillance is a moving target. ECDC maintains annual epidemiological reports and an interactive Surveillance Atlas for hantavirus data in Europe ECDC. CDC publishes weekly and annual NNDSS tables for hantavirus in the United States CDC. CDC Stacks explicitly warns that 2025 and 2026 reporting-year case counts are provisional and subject to change CDC Stacks. A useful agent should not flatten those distinctions. It should label them.
Fourth, outbreak response evolves quickly. WHO updated its hantavirus fact sheet on May 6, 2026, and emphasized integrated One Health approaches, early detection, patient care, outbreak response, and evidence-based guidance WHO. The cruise-ship cluster shows why fast, careful synthesis is useful when public attention spikes: case counts, lab confirmation, risk assessment, and operational guidance can all change within days WHO.
What the agent should not do
The danger with a topic like hantavirus is that the story almost writes itself into a thriller: invisible virus, abandoned cabins, dust in the sunlight, rare but severe disease. Good research communication has to resist the cheap version of that story.
A K-Dense Web workflow should:
- Cite primary public health sources when making clinical or prevention claims.
- Label provisional surveillance data instead of presenting it as final.
- Separate common rodent-to-human transmission from rare person-to-person transmission associated with Andes virus WHO.
- Distinguish spatial risk modeling from outbreak forecasting.
- Preserve local context, because risk in the U.S. Southwest, Scandinavia, the Balkans, East Asia, and South America is not interchangeable WHO.
That is the difference between an AI-generated explainer and a research-grade output.
The range of possibilities
For a public health team, academic lab, or science communications group, a K-Dense Web Hantavirus project can start from many different goals.
It can begin as a scientific evidence map: collect core guidance from CDC, WHO, ECDC, and national public health agencies, then build a claim table with citations, confidence levels, and unresolved questions.
It can become a surveillance analysis: extract current reporting sources, mark provisional values, generate reproducible charts, and write a methods note explaining what should be refreshed before publication.
It can become a computational workspace: build notebooks for surveillance cleaning, sequence comparison, phylogenetic summaries, spatial covariate joins, and first-pass risk models.
It can become an ecology and clinical synthesis: review rodent-host literature, climate and land-use models, outbreak case studies, diagnostic guidance, and prevention recommendations, then separate established findings from hypotheses that need field validation.
It can become a communications engine: turn the same evidence base into a public briefing, scientific memo, policy note, clinician brief, field-team guide, educator handout, figures, social graphics, and a peer-review checklist.
It can become a review environment: help humans check the claims and analyses that matter most, including case definitions, prevention advice, diagnostic nuance, sequence metadata, model assumptions, geography, policy implications, and uncertainty language.
This is not a replacement for epidemiologists, ecologists, clinicians, or public health officials. It is a way to give them a better first draft, a cleaner source trail, and more time for judgment.
Why this matters
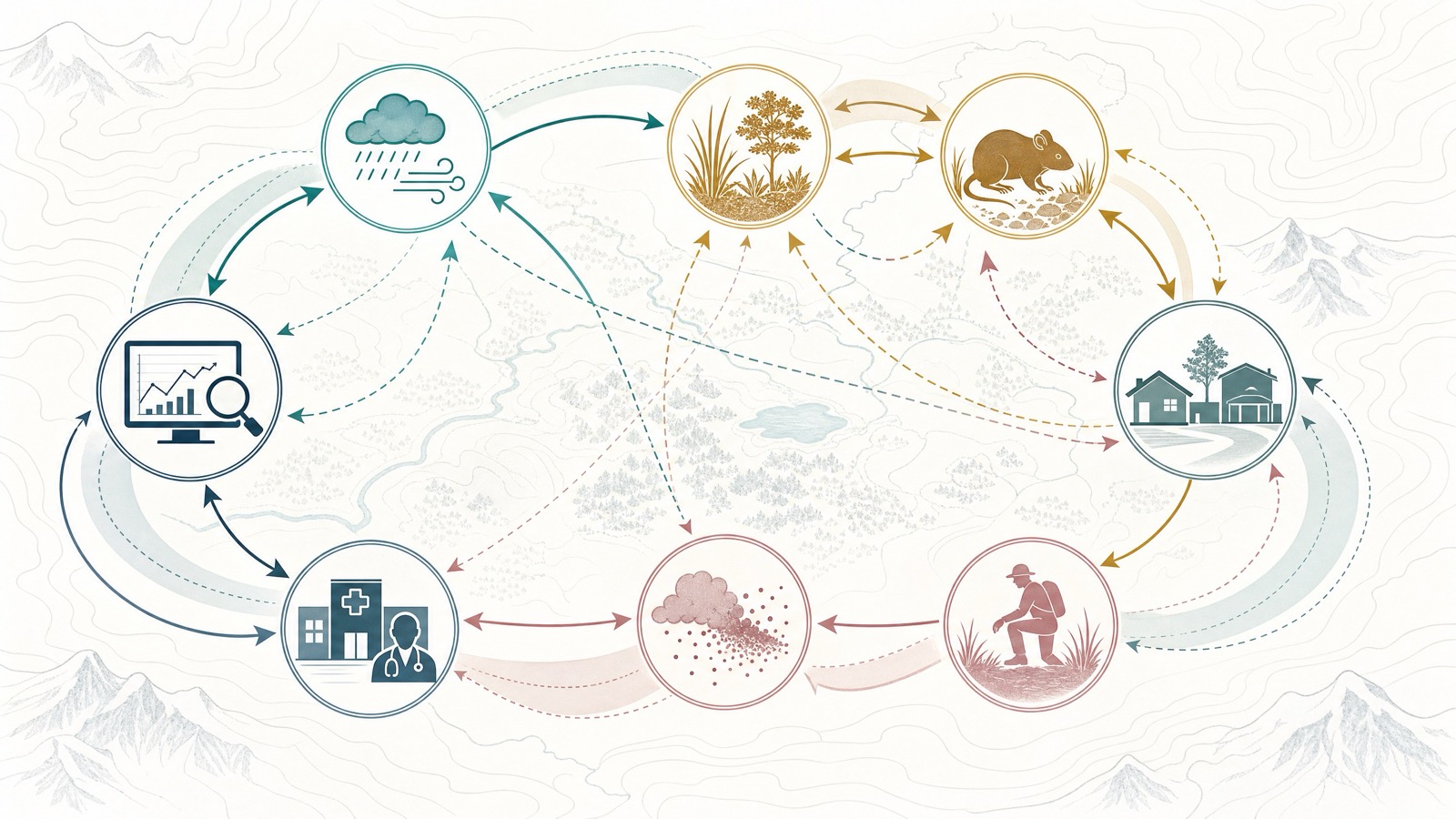
Hantavirus teaches a broader lesson about scientific AI. The hardest research problems are rarely isolated facts. They are systems. A change in rainfall can alter vegetation. Vegetation can alter rodent populations. Rodent behavior can alter human exposure. Human behavior can alter whether exposure becomes illness. Surveillance systems determine what gets seen, when it gets seen, and how confidently it can be interpreted.
K-Dense Web is useful because it can hold that whole chain in view.
For Hantavirus research, that means faster literature synthesis, clearer surveillance caveats, reproducible computational analysis, better visual communication, policy-ready briefings, and a disciplined separation between what is known, what is plausible, and what still needs fieldwork. The output is not just a blog post or a presentation. It is a reusable research object: sources, notebooks, figures, scripts, peer review, and audience-specific narratives in one place.
Hantavirus may begin with a breath of dust. Understanding it requires watching the rodents, the rain, the buildings, the clinics, and the data systems all at once.
That is exactly the kind of work K-Dense Web was made to accelerate.
Sources
- About Hantavirus, CDC
- Hantavirus Case Definition and Reporting, CDC
- CDC Stacks provisional reporting note
- Hantavirus, WHO
- Hantavirus cluster linked to cruise ship travel, WHO Disease Outbreak News
- WHO response to hantavirus cases linked to a cruise ship
- Hantavirus latest, WHO / UN Geneva
- Hantavirus-associated cluster of illness on a cruise ship, ECDC
- Surveillance and updates for hantavirus, ECDC
- Hantavirus is Associated With Open Developed Areas and Arid Climates, Wiley
